-Search query
-Search result
Showing 1 - 50 of 11,872 items for (author: ru & h)
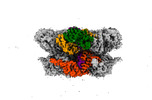
EMDB-42603: 
Human p97/VCP structure with a triazole inhibitor (NSC799462/hexamer)
Method: single particle / : Nandi P, DeVore K, Chiu PL
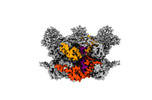
EMDB-42625: 
Human p97/VCP R155H mutant structure with a triazole inhibitor (NSC804515)
Method: single particle / : Nandi P, DeVore K, Chiu PL
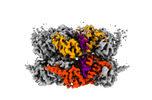
EMDB-42626: 
Human p97/VCP R155H mutant structure with a triazole inhibitor (NSC819701/up)
Method: single particle / : Nandi P, DeVore K, Chiu PL
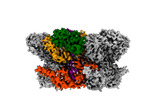
EMDB-42627: 
Human p97/VCP R155H mutant structure with a triazole inhibitor (NSC819701/down)
Method: single particle / : Nandi P, DeVore K, Chiu PL
EMDB-44748: 
Human p97/VCP structure with a triazole inhibitor (NSC799462/dodecamer)
Method: single particle / : Nandi P, DeVore K, Chiu PL

EMDB-43514: 
SPOT-RASTR - a cryo-EM specimen preparation technique that overcomes problems with preferred orientation and the air/water interface
Method: single particle / : Esfahani BG, Randolph P, Peng R, Grant T, Stroupe ME, Stagg SM
EMDB-42842: 
A Mitochondrial Replication Complex: PolG/PolG2 Bound to DNA in complex with the Single-Stranded Binding Protein (mtSSB)
Method: single particle / : Riccio AA, Bouvette J, Borgnia MJ, Copeland WC

EMDB-41766: 
CryoEM structure of D2 dopamine receptor in complex with GoA KE mutant, scFv16, and dopamine
Method: single particle / : Krumm BE, Kapolka NJ, Fay JF, Roth BL

EMDB-41776: 
CryoEM structure of D2 dopamine receptor in complex with GoA KE mutant and dopamine
Method: single particle / : Krumm BE, Kapolka NJ, Fay JF, Roth BL

EMDB-44551: 
Map of eastern equine encephalitis virus q3 spike protein in complex with VLDLR without masked refinement
Method: single particle / : Abraham J, Yang P, Li W, Fan X, Pan J

EMDB-45206: 
Reconstituted P400 Subcomplex of the human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

EMDB-45240: 
P400 subcomplex of the native human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

EMDB-45252: 
ARP module of the human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

EMDB-42291: 
Structure of the human INTS9-INTS11-BRAT1 complex
Method: single particle / : Lin M, Tong L

EMDB-42292: 
Structure of the Drosophila IntS11-CG7044(dBRAT1) complex
Method: single particle / : Lin M, Tong L

EMDB-42284: 
The structure of the Clostridium thermocellum AdhE spirosome
Method: single particle / : Ziegler SJ, Gruber JN

EMDB-37004: 
Oligonucleosome 0 DMSO
Method: subtomogram averaging / : Chen JK, Cai S, Liu T, Ruan W, Ng CT, Shi J, Surana U, Gan L

EMDB-37005: 
Oligonucleosome 9% DMSO
Method: subtomogram averaging / : Chen JK, Cai S, Liu T, Ruan W, Ng CT, Shi J, Surana U, Gan L

EMDB-38850: 
Cryo-EM structure of human dopamine transporter in apo state
Method: single particle / : Zhao Y, Li Y

EMDB-38851: 
Cryo-EM structure of human dopamine transporter in complex with dopamine
Method: single particle / : Zhao Y, Li Y

EMDB-38852: 
Cryo-EM structure of human dopamine transporter in complex with benztropine
Method: single particle / : Zhao Y, Li Y

EMDB-38853: 
structure of a proteinACryo-EM structure of human dopamine transporter in complex with GBR12909
Method: single particle / : Zhao Y, Li Y

EMDB-38854: 
Cryo-EM structure of human dopamine transporter in complex with methylphenidate
Method: single particle / : Zhao Y, Li Y

EMDB-45623: 
MicroED structure of the C11 cysteine protease clostripain
Method: electron crystallography / : Ruma YN, Bu G, Hattne J, Gonen T

PDB-9cip: 
MicroED structure of the C11 cysteine protease clostripain
Method: electron crystallography / : Ruma YN, Bu G, Hattne J, Gonen T

EMDB-42050: 
Structure of eastern equine encephalitis virus VLP in complex with VLDLR LA1
Method: single particle / : Abraham J, Yang P, Li W, Fan X, Pan J

EMDB-42055: 
Structure of eastern equine encephalitis virus VLP unliganded quasi-threefold spike protein
Method: single particle / : Abraham J, Yang P, Li W, Fan X, Pan J

EMDB-45176: 
TRRAP module of the human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

EMDB-45180: 
Second BAF53a of the human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

PDB-9c47: 
TRRAP module of the human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

PDB-9c4b: 
Second BAF53a of the human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

EMDB-41770: 
Apo form of human ATE1
Method: single particle / : Huang W, Zhang Y, Taylor DJ

EMDB-42071: 
human ATE1 in complex with Arg-tRNA and a peptide substrate
Method: single particle / : Huang W, Zhang Y, Taylor DJ

PDB-8uau: 
human ATE1 in complex with Arg-tRNA and a peptide substrate
Method: single particle / : Huang W, Zhang Y, Taylor DJ

EMDB-42948: 
Microtubule protofilament reconstruction in CCP5:microtubule class#1 complex
Method: single particle / : Chen J, Zehr EA, Gruschus JM, Szyk A, Liu Y, Tanner ME, Tjandra N, Roll-Mecak A

EMDB-44544: 
Microtubule protofilament reconstruction in CCP5:microtubule class#2 complex
Method: single particle / : Chen J, Zehr EA, Gruschus JM, Szyk A, Liu Y, Tanner ME, Tjandra N, Roll-Mecak A

EMDB-44545: 
Microtubule Protofilament reconstruction in CCP5:microtubule class#3 complex
Method: single particle / : Chen J, Zehr EA, Gruschus JM, Szyk A, Liu Y, Tanner ME, Tjandra N, Roll-Mecak A

EMDB-38300: 
Cryo-EM structure of partial dimeric WDR11-FAM91A1 complex
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D

EMDB-39863: 
Cryo-EM structure of dimeric WDR11-FAM91A1 complex
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D

EMDB-39943: 
Cryo-EM structure of dimeric WDR11-FAM91A1 complex Body2
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D

EMDB-39947: 
Cryo-EM structure of dimeric WDR11-FAM91A1 complex Body1
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D

EMDB-39949: 
Cryo-EM structure of WDR11-dm-FAM91A1 complex
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D

PDB-8xfb: 
Cryo-EM structure of partial dimeric WDR11-FAM91A1 complex
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D

PDB-8z9m: 
Cryo-EM structure of dimeric WDR11-FAM91A1 complex
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D

EMDB-16860: 
Ivabradine bound to HCN4 channel
Method: single particle / : Saponaro A, Chaves-Sanjuan A, Sharifzadeh AS, Clarke OB, Marabelli C, Bolognesi M, Thiel G, Moroni A

PDB-8ofi: 
Ivabradine bound to HCN4 channel
Method: single particle / : Saponaro A, Chaves-Sanjuan A, Sharifzadeh AS, Clarke OB, Marabelli C, Bolognesi M, Thiel G, Moroni A

EMDB-45005: 
Paired Helical Filament of tau amyloids found in Down Syndrome individuals
Method: helical / : Tse E, Ghosh U, Condello C, Southworth D

EMDB-45007: 
Straight Filament of tau amyloids found in Down Syndrome individuals
Method: helical / : Tse E, Ghosh U, Condello C, Southworth D

EMDB-45008: 
Paired Helical Filaments purified from Down Syndrome individual brain tissue applied to graphene oxide antibody affinity grids
Method: helical / : Tse E, Ghosh U, Condello C, Southworth D
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
